1. Excitation-contraction coupling in skeletal muscle is dependent on a physical interaction between the dihydropyridine receptor (DHPR) and the ryanodine receptor (RyR).
2. A number of peptides derived from the II-III loop region of the DHPR have been shown to be functionally active in stimulating the release of calcium via RyR channels. Their function has been found to correlate with the presence of a basic helical region located at the N-terminus of the II-III loop.
3. The entire recombinant skeletal DHPR II-III loop is an efficient activator of RyR1 and RyR2.
4. The skeletal DHPR II-III loop is comprised of a series of α-helices but its tertiary structure has been determined to be unstructured and flexible.
5. Fluorescence quenching experiments have been used to identify and measure the binding affinity of the II-III loop with fragments of the RyR.
In skeletal muscle, the dihydropyridine and ryanodine receptor (DHPR & RyR) are two membrane proteins expressed in the T-tubule and sarcoplasmic reticulum respectively, that play a central role in excitation-contraction coupling. EC coupling is the signal transduction process in muscle that allows a surface action potential to facilitate Ca2+ release from intracellular stores. A comprehensive description relating to the interactions between the DHPR and RyR in striated muscle have been detailed in a recent review.1 It is now widely accepted that a physical coupling between these two proteins is involved in triggering the release of Ca2+ via the RyR into the cytosol. Recent attention has focused on the exact site of interaction and a prominent site involves the loop between the second and third repeats of the skeletal DHPR α1 subunit (II-III loop).2,3 It has been shown that the II-III loop forms a physical interaction with the foot region of the RyR and any structural change in the loop is thought to be transmitted to the RyR via this interaction.4 It is therefore important to accurately determine the structural make-up of the interaction sites. From a structural viewpoint, cryoelectron microscopy has played an important role in visualising the DHPR and RyR assembly, but because of its low atomic resolution is not adequate to define structural detail at an atomic level.5,6 Our approach has been to systematically explore the structure of various segments of II-III loop by using nuclear magnetic resonance (NMR) techniques leading ultimately to the entire II-III loop. In this communication we examine the structure-function relationship of several functionally active peptide fragments ranging from the A & C peptides (as defined in 1995 by el Hayek et al.7) through to the II-III loop. In doing so we discuss how the structure of these peptides/proteins influence their interaction with specific regions in the RyR.

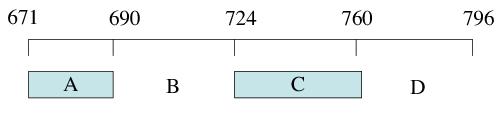
Figure 1. The II-III loop region of the skeletal DHPR. The 126 amino acid sequence has been dived into approximately 4 equal segments (A-D). The A and C regions have been shown to interact with the RyR under in vitro conditions.
In order to explore the II-III loop in closer detail it has been arbitrarily divided into four segments (A-D, Figure 1) and of these it is the A and C regions that have been the subject of intense functional and structural scrutiny. The 20 amino-acid peptide (peptide A) corresponding to the N-terminal part of the II-III loop (Thr671-Leu690) is a high-affinity activator of skeletal and cardiac RyR channels. Although in some studies the A region of the II-III loop is not required for EC coupling,8,9 it has been implicated in the overall physical interaction between the DHPR and RyR.10 A stretch of positively charged residues7,11-13 that are aligned along the surface of an α-helix structure12-14 are deemed to be an essential design feature for the activation of RyR channels by peptide A. Loss of either the positive charges or the helical structure results in a reduction in RyR activation by this peptide.13 Curiously, the scorpion toxins maurocalcine and imperatoxin A can also activate the RyR, albeit with much greater potency. Competition studies have revealed that the A segment of the II-III loop and the scorpion toxins may interact with a common binding site on the RyR.15 This is indeed plausible since it has been shown that both sets of molecules are lined with a positively charged surface.14
Peptide C (residues Glu724-Pro760) is a high affinity activator of native RyR channels at low concentration and its role in skeletal EC coupling has been well documented.16,17 It has been suggested that the C region of the II-III loop is directly responsible for the interaction with the skeletal RyR.8,9,18 Since the mechanism of EC coupling differs between skeletal and cardiac muscle, great interest surrounds whether specific modifications in the skeletal C region sequence of DHPR would lead to a loss of skeletal EC coupling. The skeletal EC coupling domain has been localized to residues 739-748 and substitution of three non-conserved residues (A739, F741 and P742) by the equivalent cardiac residues has been found to alter skeletal EC coupling.19 Using Chou-Fasman secondary structure prediction software it was predicted that one of the consequences of these amino acid substitutions would be an increase in the α-helical structure of the negatively charged residues further downstream of the mutated sites. Furthermore, it has been suggested that these structural differences could account for the non-interaction between the cardiac DHPR and RyR.19 Our recent NMR-based work provides no evidence for an increase in helical content as a result of either the proposed single or triple mutations.20 In general we find that there is some helical content between residues 740 - 749 in both the cardiac and skeletal sequences and both isoforms have common elements of secondary structure.21 The introduction of cardiac residues into the three positions 739, 741 and 742 of the skeletal DHPR II-III loop did not prevent its interaction with RyR. However, our ion channel studies do indicate that at least one residue in this region (A739) does alter the functional profile that underpins RyR and DHPR interactions.20 These findings in conjunction with the fact that the full II-III loop and the C region peptides can alter RyR activity in an isoform-independent manner suggests that in vitro, the cardiac DHPR and RyR have the potential to interact. The fact that in vivo they fail to do so may be more related to factors controlling protein expression levels and the localisation of RyR and DHPR than intrinsic differences in sequence composition and structure.
The full DHPR II-III loop has been cloned and expressed in Escherichia coli and purified using a His-tag based sequence.22 The activity of the purified protein was tested on skeletal RyR and single channel recordings performed in our laboratory show RyR activation at protein concentrations of 10 and 200 nM (as shown in Figure 2).
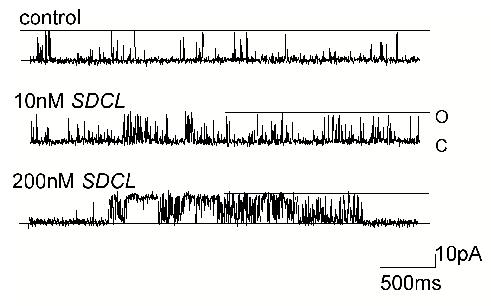
Figure 2. Single channel recordings showing activation of skeletal RyR by 10 and 200 nM recombinant skeletal II-III DHPR loop (SDCL). Activity is recorded at +40mV (mM, cis/trans)250/250 Cs+, 230/230 CH3O3S-, 20/20 Cl-, 10 mM cis Ca2+ and 1 mM trans Ca2+. Channel opening is from the closed level C to the maximum conductance (O).
NMR assignments were made with the aid of a 15N/13C double label sample using standard multinuclear NMR techniques.23 Nuclear Overhauser effects (NOEs) and dipolar measurements are used routinely in solution state NMR to define the secondary and tertiary structures of molecules. On the basis of NOE and 3J HNHA coupling data, the II-III loop secondary structure was identified as being a mixture of helical and random coil regions. However, the absence of long range NOEs indicate that the solution structure is highly disordered (Figure 3). Preliminary dynamic studies indicate that the helical regions show restricted mobility (manuscript in preparation). Confirmation of the lack of long range intramolecular interactions were observed by paramagnetic spin labelling and by residual dipolar coupling (RDC) measurements. In brief, paramagnetic spin labelling involves the introduction of paramagnetically labelled (by chemical means) residues throughout the molecule enabling the magnetic properties of atoms within 25Å of the label to be probed.24 RDCs on the other hand involve the partial alignment of molecules with the aid selected media (i.e. bacteriophage, bicelles). The magnitude of the internuclear dipolar interactions provides long-range orientational information.25 The results obtained from both of these experiments suggest the presence of a highly extended and flexible structure.
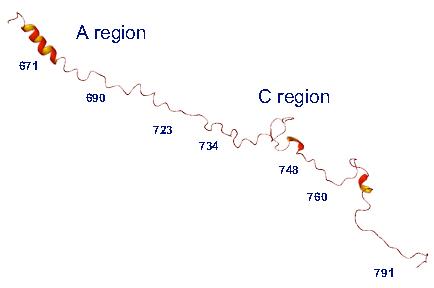
Figure 3. Solution NMR structure of the skeletal DHPR II-III loop. A possible tertiary conformation of the 126 residue recombinant DHPR II-III loop is shown. The loop does not adopt a defined tertiary fold, but is made up of a number of flexible helical segments and turns one of which is clearly seen in the A region.
In an attempt to identify the RyR II-III loop binding region, the RyR has been systematically divided into nine fragments (F1-F9); each of which has been transcribed and translated as separate protein products.26 By performing pull-down experiments with F1-F9, the location of the skeletal DHPR II-III loop binding site has been identified to lie within the F3 region (between residues 922 and 1112).26 Upon further inspection, Leong and MacLennan26 proposed that the 37-amino acid sequence between R1076 and D1112 of RyR1 was important for the binding of the II-III loop. Our goal has been to characterise the interaction between the II-III loop and the F3 fragment of the RyR. Unfortunately the F3 fragment has proven to be unstable (insoluble) at concentrations required for NMR studies. We have endeavoured to engineer shorter proteins of the F3 fragment that encompass the critical 37 amino acid region previously identified. So far the fragment containing the residues 1041-1112 (F3B) has been expressed, purified and shown to interact with the DHPR II-III loop with micromolar affinity, as measured by fluorescence quenching experiments. We are currently in the process of expanding this study with the aim of binding these RyR fragments to the DHPR II-III loop in order to map the interaction site between these two proteins. Our preliminary NMR chemical shift perturbation analyses and fluorescence studies have confirmed that the skeletal DHPR II-III loop binds weakly to the second of three SPRY domains in RyR1 (residues 1085–1208) and that the interaction sites are located within the first two α-helical regions of the DHPR II-III loop. The name SPRY arose because the domains were first identified in a Dictyostelium discoidueum splA kinase and in the mammalian RyR.27 It is becoming apparent that SPRY domains are involved in diverse biological processes including protein-protein interactions. The 2nd SPRY domain (SPRY2) is close to sequences of RyR1 located on the “clamp region” which extends towards the T-tubule and is likely to interact with the DHPR.28
The skeletal DHPR II-III loop and fragments thereof are capable of activating the RyR channel. The II-III loop protein is comprised of a series of α-helices and turns that adopt a partially folded and flexible structure. The first two α-helices, and perhaps central parts of the II-III loop are involved with a weak binding interaction with the cytosolic SPRY2 fragment of the RyR.
Our Future work will be to further explore the DHPR–RyR interaction domain by expressing sufficient quantities of stable RyR fragments capable of binding the DHPR II-III loop. The structure of the II-III loop/RyR complex will then be fully characterised using NMR techniques. This information will be important in the understanding of the physical interaction between the skeletal DHPR and RyR membrane proteins.
This work was supported by the Australian NH&MRC Project grant #22435.
1. Dulhunty AF, Haarmann CS, Green D, Laver DR, Board PG, Casarotto MG. Interactions between dihydropyridine receptors and ryanodine receptors in striated muscle. Prog. Biophys. Mol. Biol. 2002; 79: 45-75.
2. Bannister ML, Ikemoto N. Effects of peptide C corresponding to the Glu724-Pro760 region of the II-III loop of the DHP (dihydropyridine) receptor α1 subunit on the domain-switch-mediated activation of RyR1 (ryanodine receptor 1) Ca2+ channels. Biochem. J. 2006; 394: 145-52.
3. Casarotto MG, Green D, Pace S, Young J, Dulhunty AF. Activating the ryanodine receptor with dihydropyridine receptor II-III loop segments: size and charge do matter. Front. Biosci. 2004; 9: 2860-72.
4. Lu X, Xu L, Meissner G. Activation of the skeletal muscle calcium release channel by a cytoplasmic loop of the dihydropyridine receptor. J. Biol. Chem. 1994; 269: 6511-6.
5. Samso M, Wagenknecht T. Contributions of electron microscopy and single-particle techniques to the determination of the ryanodine receptor three-dimensional structure. J. Struct. Biol. 1998; 121: 172-80.
6. Samso M, Shen X, Allen PD. Structural characterization of the RyR1-FKBP12 interaction. J. Mol. Biol. 2006; 356: 917-27.
7. el-Hayek R, Antoniu B, Wang J, Hamilton SL, Ikemoto N. Identification of calcium release-triggering and blocking regions of the II-III loop of the skeletal muscle dihydropyridine receptor. J. Biol. Chem. 1995; 270: 22116-8.
8. Proenza C, Wilkens CM, Beam KG. Excitation-contraction coupling is not affected by scrambled sequence in residues 681-690 of the dihydropyridine receptor II-III loop. J. Biol. Chem. 2000; 275: 29935-7.
9. Wilkens CM, Kasielke N, Flucher BE, Beam KG, Grabner M. Excitation-contraction coupling is unaffected by drastic alteration of the sequence surrounding residues L720-L764 of the α1S II-III loop. Proc. Natl. Acad. Sci. USA 2001; 98: 5892-7.
10. Ahern CA, Bhattacharya D, Mortenson L, Coronado R. A component of excitation-contraction coupling triggered in the absence of the T671-L690 and L720-Q765 regions of the II-III loop of the dihydropyridine receptor α1s pore subunit. Biophys. J. 2001; 81: 3294-307.
11. Gurrola GB, Arevalo C, Sreekumar R, Lokuta AJ, Walker JW, Valdivia HH. Activation of ryanodine receptors by imperatoxin A and a peptide segment of the II-III loop of the dihydropyridine receptor. J. Biol. Chem. 1999; 274: 7879-86.
12. Casarotto MG, Gibson F, Pace SM, Curtis SM, Mulcair M, Dulhunty AF. A structural requirement for activation of skeletal ryanodine receptors by peptides of the dihydropyridine receptor II-III loop. J. Biol. Chem. 2000; 275: 11631-7.
13. Casarotto MG, Green D, Pace SM, Curtis SM, Dulhunty AF. Structural determinants for activation or inhibition of ryanodine receptors by basic residues in the dihydropyridine receptor II-III loop. Biophys. J. 2001; 80: 2715-26.
14. Green D, Pace S, Curtis SM, Sakowska M, Lamb GD, Dulhunty AF, Casarotto MG. The three-dimensional structural surface of two β-sheet scorpion toxins mimics that of an α-helical dihydropyridine receptor segment. Biochem. J. 2003; 370: 517-27.
15. Dulhunty AF, Curtis SM, Watson S, Cengia L, Casarotto MG. Multiple actions of imperatoxin A on ryanodine receptors: interactions with the II-III loop "A" fragment. J. Biol. Chem. 2004; 279: 11853-62.
16. Haarmann CS, Green D, Casarotto MG, Laver DR, Dulhunty AF. The random-coil 'C' fragment of the dihydropyridine receptor II-III loop can activate or inhibit native skeletal ryanodine receptors. Biochem. J. 2003; 372: 305-16.
17. Haarmann CS, Dulhunty AF, Laver DR. Regulation of skeletal ryanodine receptors by dihydropyridine receptor II-III loop C-region peptides: relief of Mg2+ inhibition. Biochem. J. 2005; 387: 429-36.
18. Tanabe T, Beam KG, Powell JA, Numa S. Restoration of excitation-contraction coupling and slow calcium current in dysgenic muscle by dihydropyridine receptor complementary DNA. Nature 1988; 336: 134-9.
19. Kugler G, Weiss RG, Flucher BE, Grabner M. Structural requirements of the dihydropyridine receptor α1S II-III loop for skeletal-type excitation-contraction coupling. J. Biol. Chem. 2004; 279: 4721-8.
20. Dulhunty AF, Karunasekara Y, Curtis SM, Harvey PJ, Board PG, Casarotto MG. Role of some unconserved residues in the "C" region of the skeletal DHPR II-III loop. Front. Biosci. 2005; 10: 1368-81.
21. Dulhunty AF, Karunasekara Y, Curtis SM, Harvey PJ, Board PG, Casarotto MG. The recombinant dihydropyridine receptor II-III loop and partly structured 'C' region peptides modify cardiac ryanodine receptor activity. Biochem. J. 2005; 385: 803-13.
22. Catanzariti AM, Soboleva TA, Jans DA, Board PG, Baker RT. An efficient system for high-level expression and easy purification of authentic recombinant proteins. Protein Sci. 2004; 13: 1331-9.
23. Cui Y, Karunasekara Y, Harvey PJ, Board PG, Dulhunty AF, Casarotto MG. 1H, 13C and 15N assignments for the II-III loop region of the skeletal dyhydropyridine receptor. J. Biomol. NMR 2005; 32: 89-90.
24. Gillespie JR, Shortle D. Characterization of long-range structure in the denatured state of staphylococcal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J. Mol. Biol. 1997; 268: 158-69.
25. Tjandra N, Bax A. Direct measurement of distances and angles in biomolecules by NMR in a dilute liquid crystalline medium. Science 1997; 278: 1111-4.
26. Leong P, MacLennan DH. A 37-amino acid sequence in the skeletal muscle ryanodine receptor interacts with the cytoplasmic loop between domains II and III in the skeletal muscle dihydropyridine receptor. J. Biol. Chem. 1998; 273: 7791-4.
27. Ponting C, Schultz J, Bork P. SPRY domains in ryanodine receptors (Ca2+-release channels). Trends Biochem. Sci. 1997; 22: 193-4.
28. Liu Z, Zhang J, Wang R, Wayne Chen SR, Wagenknecht T. Location of divergent region 2 on the three-dimensional structure of cardiac muscle ryanodine receptor/calcium release channel. J. Mol. Biol. 2004; 338: 533-45.