Fluorescence microscopy allows the non-invasive observation of biological samples in a highly sensitive and selective way. However, conventional fluorescence microscopy suffers from diffraction-limited resolution. Recent developments of super-resolution fluorescence imaging methods have found ways to bypass the diffraction barrier and enable direct optical observation of biological features approaching virtually molecular scale. An essential part of the single-molecule based localization methods such as dSTORM involves centroid fitting of serial single molecule images in order to determine the lateral localization of each fluorescent molecule with precision limited mainly by the number of emitted photons and the signal to noise ratio.
However, localization based super-resolution imaging remains challenging because of considerable hardware requirements, e.g. single molecule sensitivity, leading to expensive microscope setups. Powerful lasers, stable inverse microscope stages with high numerical aperture objectives, electron multiplied charged coupled device cameras and software cause a price of a couple of 100,000 USD for a commercial localization microscope. We have assembled an inexpensive single molecule sensitive microscope for less than 20,000 USD by questioning every component requirement. We show that the dSTORM method in combination with our open source software rapidSTORM is so robust that this rudimental microscope acquires dual color images with sub 100 nm resolution.

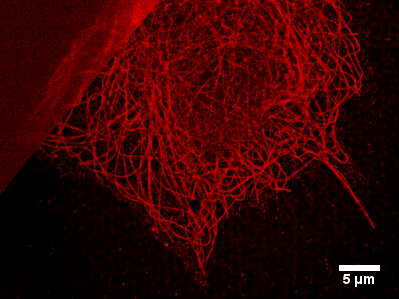
The figure shows a conventionel widefield fluorescence (upper left) and super resolution dSTORM-image (lower right) of β-tubulin labeled with Alexa Fluor 647 in fixed U2OS-cells. The image is reconstructed from a series of 15000 frames taken at a framerate of 40 frames per second, laser excitation with 640 nm and 5 kW/cm2, PBS-switching buffer with 100 mM mercaptoethylamine at a pH of 8.3.