Glutamate is the predominant excitatory neurotransmitter in the mammalian central nervous system and its extracellular levels are tightly controlled by a family of glutamate transporters also known as Excitatory Amino Acid Transporters (EAATs). The EAATs are members of a wide-ranging gene family with representatives across the spectrum of living organisms. Several X-ray crystal structures of a prokaryotic homologue of the glutamate transporter family, GltPh from Pyrococcus horikoshii, have yielded major insights into the architecture of these transporters and the conformational changes that occur during transport (Boudker et al., 2007, Reyes et al., 2009, Yernool et al., 2004).
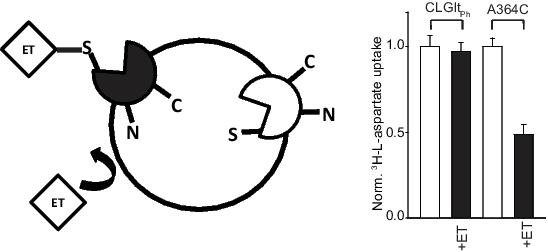
To date, the ‘open-to-inside’ structure of GltPh has not been solved but we hypothesize that a hairpin structure (HP1) opens up to allow substrate and Na+ unbinding into the cell. The aim of this study is to identify N-modified aspartate analogues that inhibit aspartate transport by GltPh and determine if they preferentially bind to the intracellular facing state of the transporter for use in structural studies. A previous study has demonstrated that purified GltPh protein reconstituted into liposomes orients randomly with ∼50% of the transporters facing right-side-out and ∼50% of the transporters facing inside-out (Ryan et al., 2009). To overcome this, we have employed a single cysteine mutant in a cysteine-less (CL) GltPh background (CLGltPhA364C) which can be modified by MTS reagents. This residue is in an extracellular facing hairpin loop (HP2) and when modified with the membrane impermeable MTS reagent MTSET, transport via the right-side-out transporters is selectively abolished allowing us to study the sidedness of GltPh functional properties as illustrated in the Figure.

We have identified six novel N-modified aspartate analogues that inhibit aspartate transport via GltPh with affinities ranging from 100-600 micromolar. We have also investigated the GltPhA364C transporter in the presence of both MTSET and the membrane permeable reagent MTSEA to probe the functional properties of the different sides of the transporter and also to determine the affinity of the inhibitors for each side of the transporter.
Boudker O, Ryan RM, Yernool D, Shimamoto K, Gouaux E. (2007) Coupling substrate and ion binding to extracellular gate of a sodium-dependent aspartate transporter. Nature 445: 387-393.
Reyes N, Ginter C, Boudker O. (2009) Transport mechanism of a bacterial homologue of glutamate transporters. Nature 462: 880-885.
Ryan RM, Compton EL, Mindell JA. (2009) Functional characterization of a Na+-dependent aspartate transporter from Pyrococcus horikoshii. Journal of Biological Chemistry 284: 17540-17548.
Yernool D, Boudker O, Jin Y, Gouaux E. (2004) Structure of a glutamate transporter homologue from Pyrococcus horikoshii. Nature 431: 811-818.