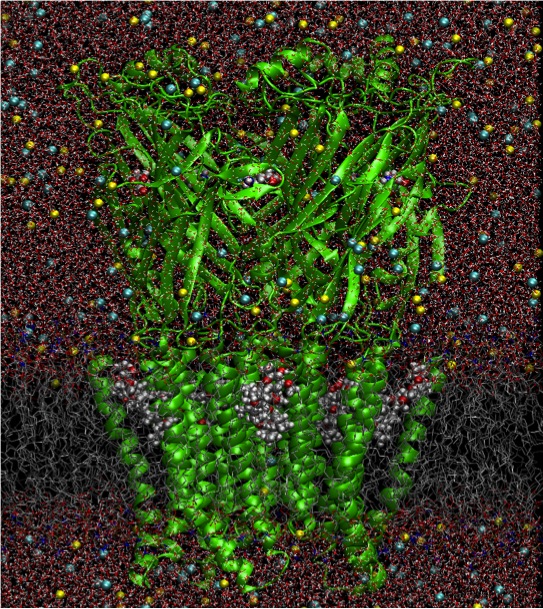
Ion channels are membrane-proteins that catalyse rapid and selective ion transport across cell membranes to enable electrical and chemical activity in the body. Recent breakthroughs in ion channel structural determination have provided a starting point from which we can begin to understand the molecular mechanisms of neuronal excitability using computer simulations. In particular, several structures of the prototypical KcsA K channel, the family of pentameric ligand-gated ion channels (pLGICs), as well as voltage-gated K and Na ion channels, have provided a glimpse of the conformational changes associated with ion channel activation/gating. We report a set of multi-microsecond simulations (approaching 0.1 millisecond in total) of fully hydrated membrane protein systems of 100,000-150,000 atoms, using the new, purpose-built DE Shaw Anton supercomputer. We observe conformational changes of GLIC and GluCl channels, and a stable putative intermediate state, that help us understand a general allosteric gating mechanism for the pLGIC family, and observe changes in the NavAb channel that shed light on voltage-gating, inactivation and permeation mechanisms. These long simulations have also revealed protein-lipid interactions that indicate specific modulating roles for different lipid components. Our studies illustrate the valuable explorative role of long atomistic computer simulations, guiding targeted and predictive computational studies of ion channel function and regulation.
