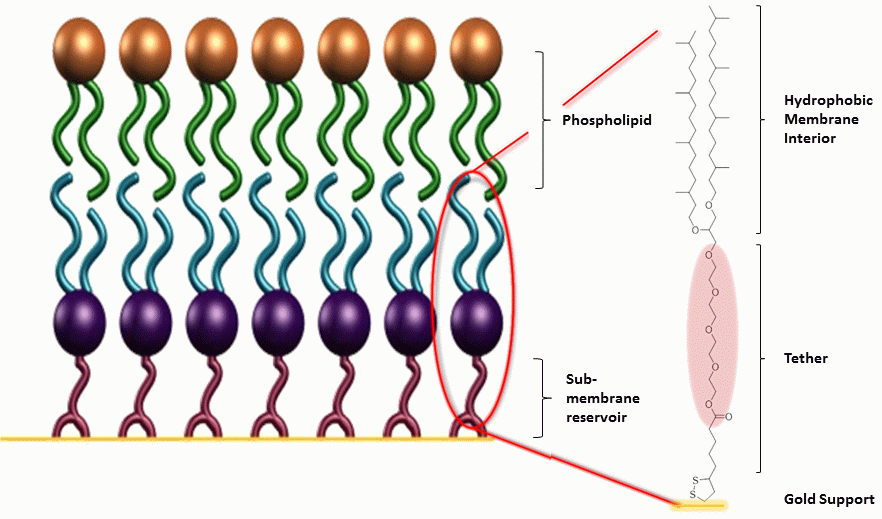
We have developed a rugged and easily assembled model of the outer membrane of Gram-negative bacteria based upon a tethered bilayer lipid membrane (tBLM) architecture. The purpose of this membrane is to enable the rapid testing of novel drug candidates designed to damage the outer membrane of a bacterial cell. This work was motivated by the significant rise of drug resistant bacterial infections (particularly Gram-negative bacteria) reported across the globe (WHO, http://www.who.int/mediacentre/factsheets/fs194/en/) and the lack of new antibiotics able to address this challenge (Lee et al., 2007).
Model membranes such as tBLMs have proven to be highly useful in a number of areas such as Alzheimer’s (Valincius et al., 2008) and HIV (Datta et al., 2011) and in biosensing applications. One of the earliest applications of tBLMs was a biosensor developed by Cornell et al., (1997). The biosensor features a bilayer membrane tethered covalently to a solid support system to construct a highly sensitive and selective biosensing device utilizing ion channel switches. The sensor is easily adaptable, as it relies on antibody-target interactions and a simple change in antibodies enables the device to be adapted to any compound of interest. A generalised schematic of a tBLM is shown in the Figure.

We have adapted this model membrane architecture to resemble the outer membrane of Gram-negative bacteria. The outer membrane of Gram-negative bacteria presents a formidable barrier to the entry of drugs into the cell, and hence is a key aspect that must be considered when developing antibiotics. Furthermore, recent studies have suggested that drug resistance appears much less likely to develop when the bacterial membrane is targeted instead of biochemical or biosynthetic pathways taking place inside the cell (Lam et al., 2016).
While the effect of membrane-damaging compounds or particles can be studied to some extent using techniques such as TEM, model membranes offer a much wider array of techniques to study the effects of novel drugs on the membrane structure. These techniques include atomic force microscopy, impedance spectroscopy and surface plasmon resonance among others. Surface Plasmon resonance enables the study of the kinetics of drug-membrane interactions and neutron scattering enables the measurement of sub-nanometer structural changes induced in the membrane by antibiotic compounds. A detailed understanding of the mechanism by which an antibiotic compound interacts with the bacterial membrane is essential if we are to develop more effective antibiotics in the future.
Cornell BA, Braach-Maksvytis VL, King LG, Osman PD, Raguse B, Wieczorek L, Pace RJ. (1997) A biosensor that uses ion-channel switches. Nature 387, 580-3.
Datta SAK, Heinrich F, Raghunandan S, Kruege S, Curtis JE, Rein A, Nanda H. (2011) HIV-1 Gag extension: conformational changes require simultaneous interaction with membrane and nucleic acid. J Mol Biol 406, 205-214.
Lam SJ, O'Brien-Simpson NM, Pantarat N, Sulistio A, Wong EH, Chen YY, Lenzo JC, Holden JA, Blencowe A, Reynolds EC, Qiao GG. (2016) Combating multidrug-resistant Gram-negative bacteria with structurally nanoengineered antimicrobial peptide polymers. Nat Microbiol 1, 16162.
Lee JH, Jeong SH, Cha S-S, Lee SH. (2007) A lack of drugs for antibiotic-resistant gram-negative bacteria. Nat Rev Drug Disc 6, 938-939.
Valincius G, Heinrich F, Budvytyte R, Vanderah DJ, McGillivray DJ, Sokolov Y, Hall JE, Lösche M. (2008) Soluble amyloid β-oligomers affect dielectric membrane properties by bilayer insertion and domain formation: implications for cell toxicity. Biophys J 95, 4845-4861.